About
What is a "hetnet"?
A network (also known as a graph) is a conceptual representation of a group of things — called nodes — and the relationships between them — called edges. Typically, a network has only one type of node and one type of edge. But in many cases, it is necessary to be able to distinguish between different types of entities and relationships.
A hetnet (short for heterogeneous information network) is a network where nodes and edges can be multiple types. This additional dimension allows a hetnet to accurately describe more complex data. Hetnets are particularly useful in biomedicine, where it is important to capture the conceptual distinctions between various components and mechanisms, such as genes and diseases, or upregulation and binding.
The prefix meta is used on this site to refer to the type of the node/edge (e.g. compound), as opposed to the specific node/edge itself (e.g. acetaminophen).
What is Hetionet?
Hetionet is a hetnet of biomedical knowledge. It encodes relationships uncovered by millions of studies conducted over the last half-century into a single resource. The network is constructed from a collection of publicly available databases, and is itself open-source and free to use, barring any upstream restrictions.
Hetionet enables scientists and biologists to formulate novel hypotheses, predictions, and other valuable insights by connecting an existing body of biomedical data across multiple levels and types in a convenient, accessible, holistic way.
Why was Hetionet made?
Hetionet was originally created as part of Project Rephetio, a study that utilized the benefits of hetnets to predict new uses for existing drugs. Although the original resources for the network were selected for drug repurposing, Hetionet is now useful in a much broader sense, and has been used for a variety of purposes.
Hetionet was also made to alleviate some of the inaccuracies and inconveniences of using other integrative networks, or trying to use multiple, separate databases in the same analysis. It unifies data from several different, disparate sources into a single, comprehensive, accessible, common-format network.
What's in Hetionet?
Hetionet combines information from 29 public databases. The network contains 47,031 nodes of 11 types and 2,250,197 edges of 24 types.
| Metanode | Description |
|---|---|
| Gene | Protein-coding human genes. From Entrez Gene. |
| Compound | Approved small molecule compounds with documented chemical structures. From DrugBank. |
| Anatomy | Anatomical structures, excluding structures that are known not to be found in humans. From Uberon. |
| Disease | Complex diseases, selected to be distinct and specific enough to be clinically relevant yet general enough to be well annotated. From Disease Ontology. |
| Symptom | Signs and Symptoms (i.e. clinical abnormalities that can indicate a medical condition). From the MeSH ontology. |
| Side Effect | Adverse drug reactions. From SIDER/UMLS. |
| Biological Process | Larger processes or biological programs accomplished by multiple molecular activities. From Gene Ontology. |
| Cellular Component | The locations relative to cellular structures in which a gene product performs a function. From Gene Ontology. |
| Molecular Function | Activities that occur at the molecular level, such as "catalysis" or "transport". From Gene Ontology. |
| Pathway | A series of actions among molecules in a cell that leads to a certain product or change in the cell. From WikiPathways, Reactome, and Pathway Interaction Database. |
| Pharma |
"Chemical/ |
The metagraph diagram below illustrates the connectivity between different types of nodes and edges in the network.
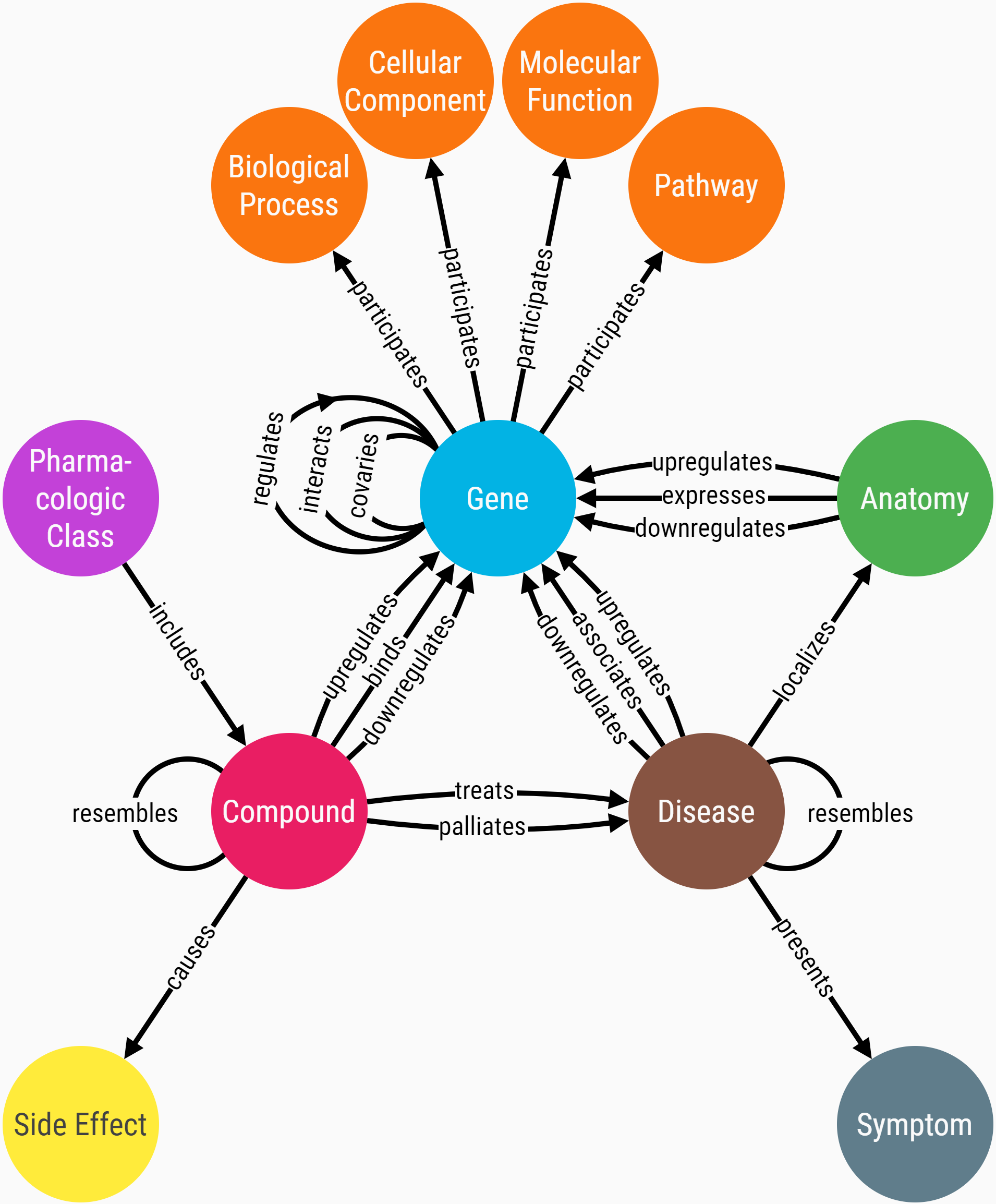
Notice that multiple types of edges can connect the same two nodes. For example, a compound can have a relationship to a gene by binding its protein and/or by upregulating it. Since the biological effects of these two mechanisms are very different, they are given separate representations in the network.
How was Hetionet made?
Hetionet was created openly, with a process of continuous public discussion and feedback. In general, great care was taken to "clean" and "de-duplicate" the network such that it would be as easy and effective to use as possible.
A standardized vocabulary was selected to define each metanode (e.g. Entrez Gene for genes or Disease Ontology for diseases). These vocabularies provide a robust, common backbone to overlay nodes and edges onto without repeating equivalent concepts.
Edges were extracted from omics-scale resources (ranging from text-mining to high-throughput screens) and their identifiers were converted to the appropriate node vocabularies. Edges were consolidated such that one relationship is encoded into a single edge, even when multiple studies or resources have reported it.
See here for a more detailed description of the methodologies used to construct the network.
Acknowledgements
Hetionet was originally developed by Daniel Himmelstein and collaborators in the Baranzini Lab at the University of California, San Fransisco, as part of Project Rephetio. Thank you to the team of online contributors who provided feedback throughout the course of the project.
Other applications of Hetionet and algorithms for hetnets continue to be developed by Daniel in the Greene Lab at the University of Pennsylvania.
Daniel's work in the Baranzini Lab was supported by the National Science Foundation Graduate Research Fellowship (1144247). Sergio's work was in part supported by the National Institutes of Health (R01 NS088155). Development of hetnet software, applications, and algorithms in the Greene Lab is supported by the Gordon and Betty Moore Foundation (GBMF4552) and through a research collaboration with Pfizer Worldwide Research and Development. For a comprehensive list of contributors, grants, and funding sources, see the acknowledgement sections of related publications.
Cite Drug Repurposing (Project Rephetio)
For the topics eLife: Project Rephetio, drug repurposing, Hetionet v1.0, data licensing, realtime open science, consensus LINCS L1000 signatures, hetnets, HNEP, cite the following:
Himmelstein DS, Lizee A, Hessler C, Brueggeman L, Chen SL, Hadley D, Green A, Khankhanian P, Baranzini SE
eLife (2017)
DOI: 10.7554/elife.26726 · PMID: 28936969 · PMCID: PMC5640425
Cite Prioritizing Disease-Associated Genes
For the topics PLOS: DWPC, predicting disease-gene associations, hetnet permutation, HNEP, cite the following:
Himmelstein DS, Baranzini SE
PLOS Computational Biology (2015)
DOI: 10.1371/journal.pcbi.1004259 · PMID: 26158728 · PMCID: PMC4497619
Contact
For questions or feedback regarding existing projects, please open an issue on the relevant GitHub repository (under @hetio or @dhimmel). For other inquiries, send an email to daniel.himmelstein@gmail.com.